In a paper published in Nature today, titled “The genome of a Late Pleistocene human from a Clovis burial site in western Montana,” by Rasmussen et al, the authors conclude that the DNA of a Clovis child is ancestral to Native Americans. Said another way, this Clovis child was a descendant, along with Native people today, of the original migrants from Asia who crossed the Bering Strait.
This paper, over 50 pages including supplemental material, is behind a paywall but it is very worthwhile for anyone who is specifically interested in either Native American or ancient burials. This paper is full of graphics and extremely interesting for a number of reasons.
First, it marks what I hope is perhaps a spirit of cooperation between genetic research and several Native tribes.
Second, it utilized new techniques to provide details about the individual and who in world populations today they most resemble.
Third, it utilized full genome sequencing and the analysis is extremely thorough.
Let’s talk about these findings in more detail, concentrating on information provided within the paper.
The Clovis are defined as the oldest widespread complex in North America dating from about 13,000 to 12,600 calendar years before present. The Clovis culture is often characterized by the distinctive Clovis style projectile point. Until this paper, the origins and genetic legacy of the Clovis people have been debated.
about 13,000 to 12,600 calendar years before present. The Clovis culture is often characterized by the distinctive Clovis style projectile point. Until this paper, the origins and genetic legacy of the Clovis people have been debated.
These remains were recovered from the only known Clovis site that is both archaeological and funerary, the Anzick site, on private land in western Montana. Therefore, the NAGPRA Act does not apply to these remains, but the authors of the paper were very careful to work with a number of Native American tribes in the region in the process of the scientific research. Sarah L. Anzick, a geneticist and one of the authors of the paper, is a member of the Anzick family whose land the remains were found upon. The tribes did not object to the research but have requested to rebury the bones.
The bones found were those of a male infant child and were located directly below the Clovis materials and covered in red ochre. They have been dated to about 12,707-12,556 years of age and are the oldest North or South American remains to be genetically sequenced.
All 4 types of DNA were recovered from bone fragment shavings: mitochondrial, Y chromosome, autosomal and X chromosome.
Mitochondrial DNA
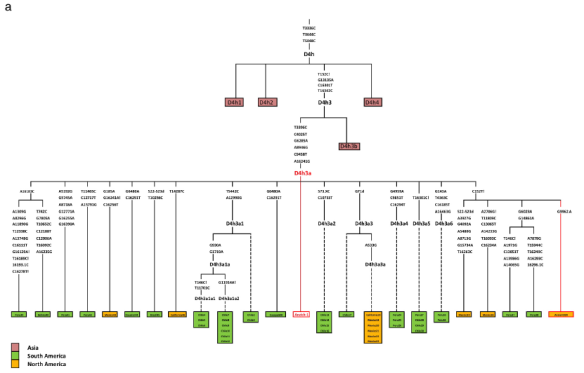
The mitochondrial haplogroup of the child was D4h3a, a rather rare Native American haplogroup. Today, subgroups exist, but this D4h3a sample has none of those mutations so has been placed at the base of the D4h3a tree branch, as shown below in a grapic from the paper. Therefore, D4h3a itself must be older than this skeleton, and they estimate the age of D4h3a to be 13.000 plus or minus 2,600 years, or older.
Today D4h3a is found along the Pacific coast in both North and South America (Chile, Peru, Ecuador, Bolivia, Brazil) and has been found in ancient populations. The highest percentage of D4h3a is found at 22% of the Cayapa population in Equador. An ancient sample has been found in British Columbia, along with current members of the Metlakatla First Nation Community near Prince Rupert, BC.
Much younger remains have been found in Tierra del Fuego in South America, dating from 100-400 years ago and from the Klunk Mound cemetery site in West-Central Illinois dating from 1800 years ago.
It’s sister branch, D4h3b consists of only one D4h3 lineage found in Eastern China.
Y Chromosomal DNA
The Y chromosome was determined to be haplogroup Q-L54. Haplogroup Q and subgroup Q-L54 originated in Asia and two Q-L54 descendants predominate in the Americas: Q-M3 which has been observed exclusively in Native-Americans and Northeastern Siberians and Q-L54.
The tree researchers constructed is shown below.
They estimate the divergence between haplogroups Q-L54 and Q-M3, the two major haplogroup Q Native lines, to be about 16,900 years ago, or from between 13,000 – 19,700.
The researchers shared with us the methodology they used to determine when their most common recent ancestor (MCRA) lived.
“The modern samples have accumulated an average of 48.7 transversions [basic mutations] since their MCRA lived and we observed 12 in Anzick. We infer an average of approximately 36.7 (48.7-12) transversions to have accumulated in the past 12.6 thousands years and therefore estimate the divergence time of Q-M3 and Q-L54 to be approximately 16.8 thousands years (12.6ky x 48.7/36.7).”
Autosomal
They termed their autosomal analysis “genome-wide genetic affinity.” They compared the Anzick individual with 52 Native populations for which known European and African genetic segments have been “masked,” or excluded. This analysis showed that the Anzick individual showed a closer affinity to all 52 Native American populations than to any extant or ancient Eurasian population using several different, and some innovative and new, analysis techniques.
Surprisingly, the Anzick infant showed less shared genetic history with 7 northern Native American tribes from Canada and the Artic including 3 Northern Amerind-speaking groups. Those 7 most distant groups are: Aleutians, East Greenlanders, West Greenlanders, Chipewyan, Algonquin, Cree and Ojibwa.
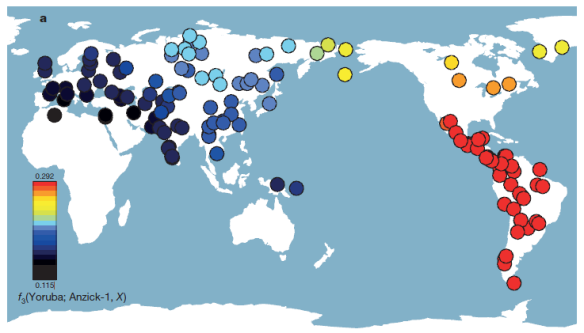
They were closer to 44 Native populations from Central and South America, shown on the map below by the red dots. In fact, South American populations all share a closer genetic affinity with the Anzick individual than they do with modern day North American Native American individuals.
The researchers proposed three migration models that might be plausible to support these findings, and utilized different types of analysis to eliminate two of the three. The resulting analysis suggests that the split between the North and South American lines happened either before or at the time the Anzick individual lived, and the Anzick individual falls into the South American group, not the North American group. In other words, the structural split pre-dates the Anzick child. They conclude on this matter that “the North American and South American groups became isolated with little or no gene flow between the two groups following the death of the Anzick individual.” This model also implies an early divergence between these two groups.
In Eurasia, genetic affinity with the Anzick individual decreases with distance from the Bering Strait.
The researchers then utilized the genetic sequence of the 24,000 year old MA-1 individual from Mal’ta, Siberia, a 40,000 year old individual “Tianyuan” from China and the 4000 year old Saqqaq Palaeo-Eskimo from Greenland.
Again, the Anzick child showed a closer genetic affinity to all Native groups than to either MA-1 or the Saqqaq individual. The Saqqaq individual is closest to the Greenland Inuit populations and the Siberian populations close to the Bering Strait. Compared to MA-1, Anzick is closer to both East Asian and Native American populations, while MA-1 is closer to European populations. This is consistent with earlier conclusions stating that “the Native American lineage absorbed gene flow from an East Asian lineage as well as a lineage related to the MA-1 individual.” They also found that Anzick is closer to the Native population and the East Asian population than to the Tianyuan individual who seems equally related to a geographically wide range of Eurasian populations. For additional information, you can see their charts in figure 5 in their supplementary data file.
I have constructed the table below to summarize who matches who, generally speaking.
In addition, a French population was compared and only showed an affiliation with the Mal’ta individual and generically, Tianyuan who matches all Eurasians at some level.
Conclusions
The researchers concluded that the Clovis infant belonged to a meta-population from which many contemporary Native Americans are descended and is closely related to all indigenous American populations. In essence, contemporary Native Americans are “effectively direct descendants of the people who made and used Clovis tools and buried this child,” covering it with red ochre.
Furthermore, the data refutes the possibility that Clovis originated via a European, Solutrean, migration to the Americas.
I would certainly be interested to see this same type of analysis performed on remains from the eastern Canadian or eastern seaboard United States on the earliest burials. Pre-contact European admixture has been a hotly contested question, especially in the Hudson Bay region, for a very long time, but we have yet to see any pre-Columbus era contact burials that produce any genetic evidence of such.
Additionally, the Ohio burial suggests that perhaps the mitochondrial DNA haplogroup is or was more widespread geographically in North American than is known today. A wider comparison to Native American DNA would be beneficial, were it possible. A quick look at various Native DNA and haplogroup projects at Family Tree DNA doesn’t show this haplogroup in locations outside of the ones discussed here. Haplogroup Q, of course, is ubiquitous in the Native population.
National Geographic article about this revelation including photos of where the remains were found. They can make a tuft of grass look great!
Another article can be found at Voice of America News.
Science has a bit more.
If you’d like to take a DNA test, click here.




Just a side note: my great grandmother was an adopted Native American. We know this from family history, but have no proof. Our dentist upon examining my daughter’s teeth and x-rays asked: ” Are you Japanese?” We said “part Native American. He was astounded. Thanks for the article.